Faculty
|

|
Please click here for more information.
|
Visiting Faculty
|
|
|
Youzhi Tang
|
|
|
Kazuma Yasuhara
I am interested in the molecular design of membrane-active molecules based on the physicochemical understanding of their interaction mechanism. Also, I study biological application of supramolecular assemblies such as lipid nanodiscs formed with synthetic lipids.
|
Senior Scientists
|

|
Hyunil Jo (C.V.)
My research area spans from Medicinal Chemistry to Protein Engineering. I have been working on developments of antimalarial, anti-HIV, Integrin inhibitors, Splicing corrector for SMA (Spinal Muscular Atrophy) treatment, and M2 proton channel inhibitor. For protein engineering, protein structure restriction by side chain modification, installation of IR and UV-resonance Raman probe have been intensively studied.
|

|
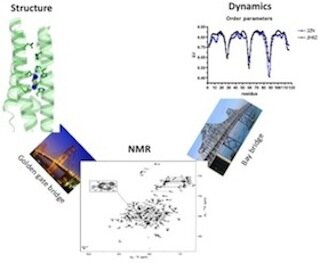
Yibing Wu (C.V.)
My research in general focuses on structure and dynamic study of protein using multidimensional NMR spectroscopy. At different Zn concentrations, I have recently solved two Diiron structures, which help us make new designs to stabilize the structure and optimize the active site. Structural study on AM2 protein is another ongoing project. Current focus is to identify good inhibitors and obtain high-resolution solution structure of M2 S31N in presence of drug to investigate the detail binding sites.
|
Postdocs
|

|
Manasi Bhate ()
I am interested developing quantitative mechanistic models that describe the transmission of information across the cellular membrane. A living cell is separated from its environment by a cellular membrane and much of life depends on the accurate and efficient signal transduction across this membrane. I use a combination of protein design, quantitative biophysics and NMR spectroscopy to study the structure and dynamics of transmembrane proteins that mediate signal transduction.
|
|
|
Billy Egan
Neurodegenerative diseases such as Alzheimer's, Parkinsons and Creutzfeldt-Jakob Disease (CJD) are associated with aggregation of fibrillar deposits in brain tissue. The onset of -sheet polymerization of Amyloid- (A) peptides is believed to play a central role in the pathogenesis of these neurodegenerative plaques. I am performing the synthesis and purification of A prions, which are capable of undergoing fibrillization in aggregation kinetics studies.
Included in my research is the preparation of A mutant variants with isotope labeled 13C and 15N residues at multiple selected sites on the peptide, which has applications in a solids NMR (SSNMR) study. This will probe the structure and provide a possible explanation for the observed change in morphology and stability of A1-40. This also will allow site-specific assignment by inspection of the 2D 13C-13C 15N-15N correlation spectra, and enable us to examine the extent in which the labeled residues are ordered and to determine their secondary structure. The sidechains of selected residues will also be labeled with 13C=18O to probe hydration and the strength of hydrogen-bonding as A40 and A42 progress from oligomers to fibrils.
|

|
Nate Joh (C.V.) ()
Transmembrane transport, central to many vital cell functions including nutrient uptake, signaling and drug efflux, is mediated by various transporters. To learn the transporter function and mechanism, I take both "top-down" and "bottom-up" approaches.
In a top-down approach where I learn by examining the natural system, proton conduction by matrix protein M2 from influenza A virus is investigated, as well as its neutralization by anti-M2 antibodies, using various biophysical methods including crystallography.
Alternatively, in a bottom-up approach, I apply the first principles to generate the transport functionality through engineering the alternate-access mechanism using de-novo protein design method.
|
|
|
Thomas Lemmin
The self-assembly of prion precursors into oligomers and fibers has been linked to several neurodegenerative and systemic disorders. In particular, the aggregation of Amyloid- (A) prions and the formation of extracellular A plaques in brain tissue have been associated with the neuropathological process of Alzheimer’s disease (AD). I am interested in integrating molecular modeling techniques and experimental data to determine the oligomeric and polymeric species of the A sequence at different points during its self-assembly process. These models will permit us to design small molecules, peptides and peptide mimetics that modulate the self-assembly process.
|
|
|
Reza Malmirchegini
Many relevant drug targets are membrane proteins that contain transmembrane (TM) helices. I am interested in TM helices and their interaction with one another. I am working on developing a high-throughput genetic system that will allow detection of both homo- and heterooligomerization of TM helices in mammalian cells by utilizing FRET (Förster Resonance Energy Transfer). My interests are therefore two-fold: the first is a proteomics approach to use this system to identify all TM helix partners in a given organism. The second is a pharmacological approach in which this system will be used to rapidly screen through a large library of potential TM peptide inhibitors. I am also interested in the TM region present in the HIV coat protein gp41. I am working on using directed evolution on a chimeric protein containing the MPER (membrane proximal external region) that will elicit production of broadly neutralizing antibodies.
|

|
Michelle McCully ()
This lab is developing a vaccine for HIV using the membrane proximal external region (MPER) of the HIV-1 envelope glycoprotein. Broadly neutralizing monoclonal antibodies that target the MPER exist in chronically infected individuals, and these antibodies provide protection against infection in macaques. Currently, there is little structural information available on the MPER, so I am working towards characterizing the MPER using a combination of molecular dynamics simulations and experimental techniques. These data will be used in rational vaccine design to stabilize the antibody-eliciting conformation of the MPER.
|

|
Mimi Nick
Alzheimer's Disease is characterized by the accumulation of pathogenic protein aggregates that share a specific fibrillar conformation called amyloid. While amyloid fibrils are known to be very stable and contribute to proteotoxicity, the soluble oligomeric complexes that precede fibril formation may be even more cytotoxic. However, very little is known about the molecular structure of these transient conformers. I am interested in creating stable oligomers that will allow us to elucidate the protein structure of these typically fleeting accumulations and determine how specific conformations contribute to neurodegeneration.
|
|
|
Nathan Schmidt
|

|
Gözde Ulas
My research involves the design of small peptides in order to modulate activity of target protein complexes. By stabilizing key intermediates of target protein function, we aim to capture transient stages of proteins of interest. One intermediary stage that we would like to stabilize is the intermediary fusion stage of HIV-1 gp41 envelope glycoprotein.
|

|
Shaoqing Zhang
My current projects focus on the relationship between the sequence and conformation of transmembrane helices using structural bioinformatic approaches. I am also working on the structural and functional analysis of transmembrane proteins in signal transduction, specifically in the Tpo/TpoR system.
|
Grduate Students
|
|
|
Leo Gendelev
Designed proteins that can modulate protein-protein interactions have an advantage over small molecules when it comes to flat protein-protein interfaces. To this end I am using a nature-inspired approach involving both computational methods and directed evolution techniques such as phage display to target a key interaction in the activation of the family of EGFR Kinases. The C-lobe of one EGFR Kinase can allosterically activate the N-lobe of a partner EGFR Kinase, so inhibiting this interaction is a promising approach to treat various types of cancers. Computationally I am mining the protein data bank for motifs based on nature-preferred interaction geometries to use as a starting point, and then scanning for protein-based scaffolds that can accommodate these motifs. I will validate as well as affinity mature the most promising designs using phage display, with the eventual goal of developing a protein-based therapeutic.
|

|
Bruk Mensa
My research focuses on the bactericidal mechanism of action of small molecule antimicrobial peptide mimetics. I also study bacterial signal transduction via two-component systems, particularly focusing on signal propagation mechanisms of virulence-associated sensor kinases and solving atomic-level structures of full-length sensor kinases.
|

|
Kathleen Molnar
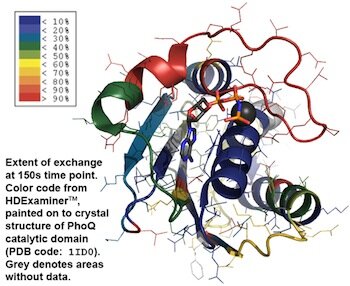
Mass spectrometry is critical to many quantitative biophysical techniques. One such technique, hydrogen/deuterium exchange is used to measure the dynamics of proteins in vitro. Pictured below are the exchange dynamics of the catalytic domain of PhoQ. My research focuses this bacterial two-component system PhoQ/P and determining the protein dynamics during signaling.
|

|
Jessica Thomaston
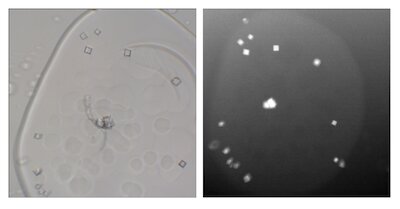
My primary interests are membrane protein structure and function. Currently, the focus of my efforts is a structural study of the AM2 and BM2 proton channels of influenza. Influenza AM2 was a drug target before viral resistance developed in the past decade, and the BM2 channel has no known ligands. I use lipidic cubic phase (LCP) techniques in an attempt to crystallize these transmembrane channels in a physiologically relevant environment.
|
Staff
|

|
Lili Li
Administrative Assistant for the lab
|

















